Reposted from microBEnet… Building science measurements in the Hospital Microbiome Project: Part 1
By Brent Stephens on November 6, 2013
This post was originally written for microBEnet and is copied here.
First of all, Happy Halloween everyone. I think my costume this year will be a blogger!
For those that don’t know me, I’m Brent Stephens, an assistant professor in the Department of Civil, Architectural and Environmental Engineering at Illinois Institute of Technology in Chicago, IL. I call my research team the Built Environment Research Group and my teaching and research interests are primarily in characterizing energy and air quality in buildings. Some people would also call me a “building scientist,” and it’s that description that brought me to microBEnet. Since Fall 2012, my team and I have been working with Jeff Siegel at the University of Toronto on Jack Gilbert’s Sloan-funded Hospital Microbiome Project.
The Hospital Microbiome Project is huge in scope: nearly 13,000 microbial samples are being taken from a variety of surfaces in a brand new hospital pavilion at the University of Chicago. Fortunately, microbial sampling began about a month before the hospital actually opened up to patients, and has continued for nearly a year since then (we’re about 9 months into the project). This allows Jack’s team to tracking how microbial communities evolve over time in a critical hospital environment as patients occupy the space. Microbial samples are being taken within 10 patient rooms in this new hospital (5 on one floor and 5 on another) from both patient bodies and surfaces such as floors, bed rails, faucet handles, as well as surfaces such as computer mice, chairs, and countertops at a central nurse’s station on each floor.
This all sounds like an amazing study (and it is), but where do we fit in as building scientists??
 Well, we’re there in the new hospital too. But instead of roving around the hospital taking swabs, we snuck in just before the hospital opened and installed a variety of sensors to track a wealth of building environmental and operational data over the course of the project. As Elizabeth Glass and coauthors recently described, there are a number of factors that can influence microbial abundance and diversity in buildings, including human occupancy, temperature, and relative humidity. Airflow is another important factor, including whether rooms are pressurized or depressurized relative to adjacent spaces and outdoor air ventilation rates. Higher ventilation rates can dilute concentrations of indoor-generated bioaerosols, which may in turn impact the amount deposited onto surfaces. Many buildings (including this hospital) will vary ventilation rates depending on outdoor temperature by using a mechanical control system called an economizer. So we’re doing our best to measure these variables in the hospital. Hopefully at the end of the project, we’ll have a robust, long-term data set of building environmental and operational parameters that we can use to explore correlations with microbial abundance and/or diversity in the hospital. Many of these parameters are listed in the Built Environment minimal metadata package led by Elizabeth Glass and Lynn Schriml.
Well, we’re there in the new hospital too. But instead of roving around the hospital taking swabs, we snuck in just before the hospital opened and installed a variety of sensors to track a wealth of building environmental and operational data over the course of the project. As Elizabeth Glass and coauthors recently described, there are a number of factors that can influence microbial abundance and diversity in buildings, including human occupancy, temperature, and relative humidity. Airflow is another important factor, including whether rooms are pressurized or depressurized relative to adjacent spaces and outdoor air ventilation rates. Higher ventilation rates can dilute concentrations of indoor-generated bioaerosols, which may in turn impact the amount deposited onto surfaces. Many buildings (including this hospital) will vary ventilation rates depending on outdoor temperature by using a mechanical control system called an economizer. So we’re doing our best to measure these variables in the hospital. Hopefully at the end of the project, we’ll have a robust, long-term data set of building environmental and operational parameters that we can use to explore correlations with microbial abundance and/or diversity in the hospital. Many of these parameters are listed in the Built Environment minimal metadata package led by Elizabeth Glass and Lynn Schriml.
In the rest of this post I’d like to further describe the variables we’re measuring and how we’re doing it. We’ll save most of our results for a future post.
In each of the sample patient rooms of the hospital we’re measuring the following building science parameters:
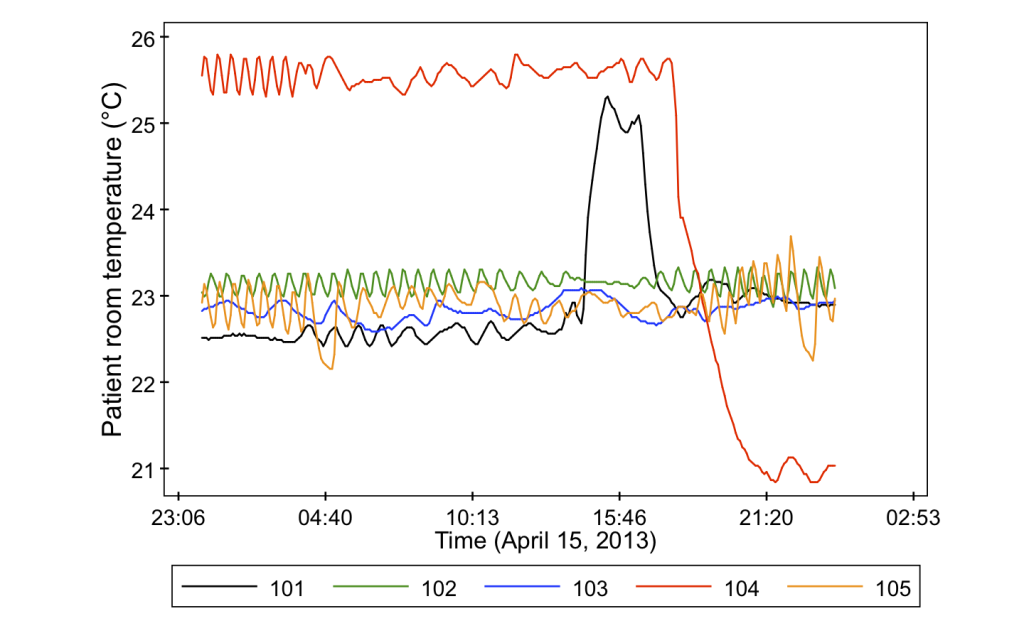
- Indoor air temperature in a central location (also at the common nurse stations)
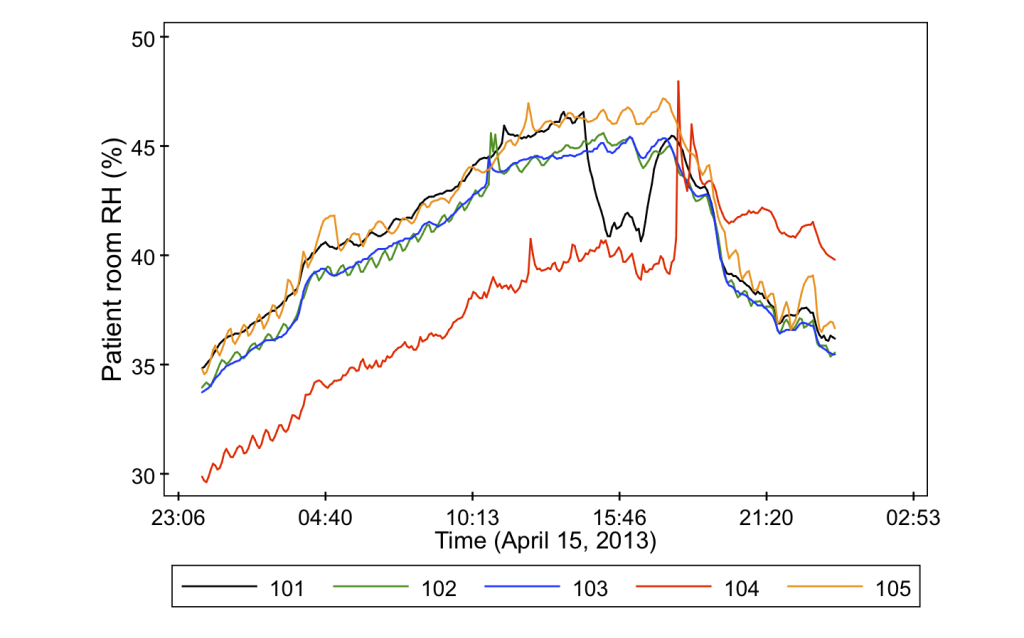
- Indoor air relative humidity in a central location (also at the common nurse stations)
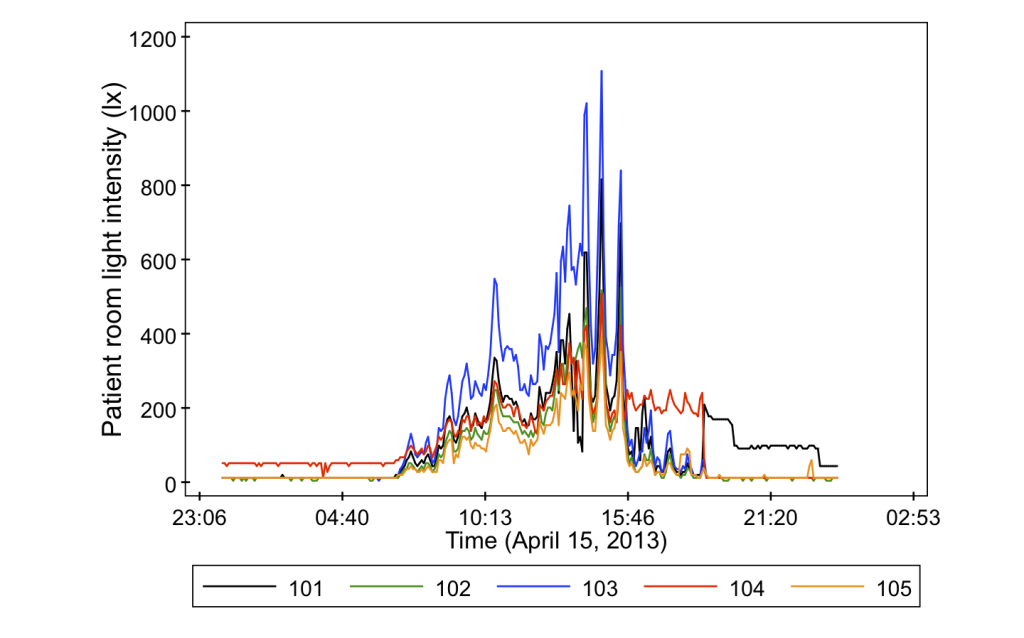
- Light intensity in a central location (also at the common nurse stations)
- Indoor CO2 concentrations (as a surrogate for occupancy and a check on ventilation rates)
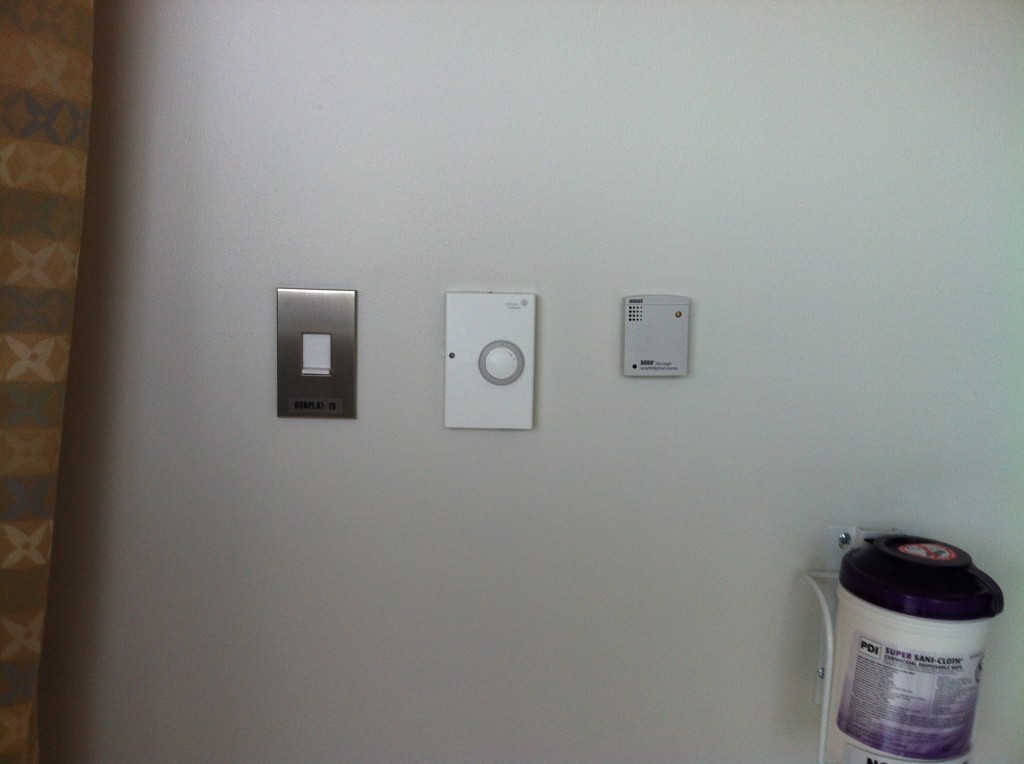
- Room pressurization relative to the hallway
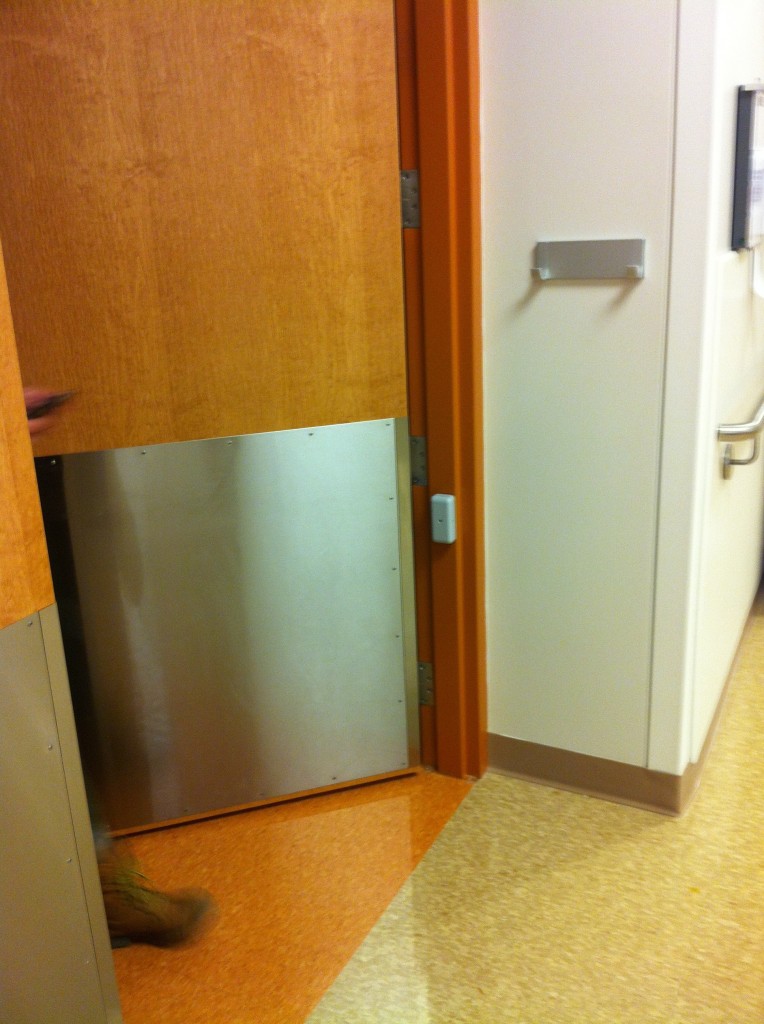
- Human occupancy by an infrared beam break sensor installed at the patient room doorway
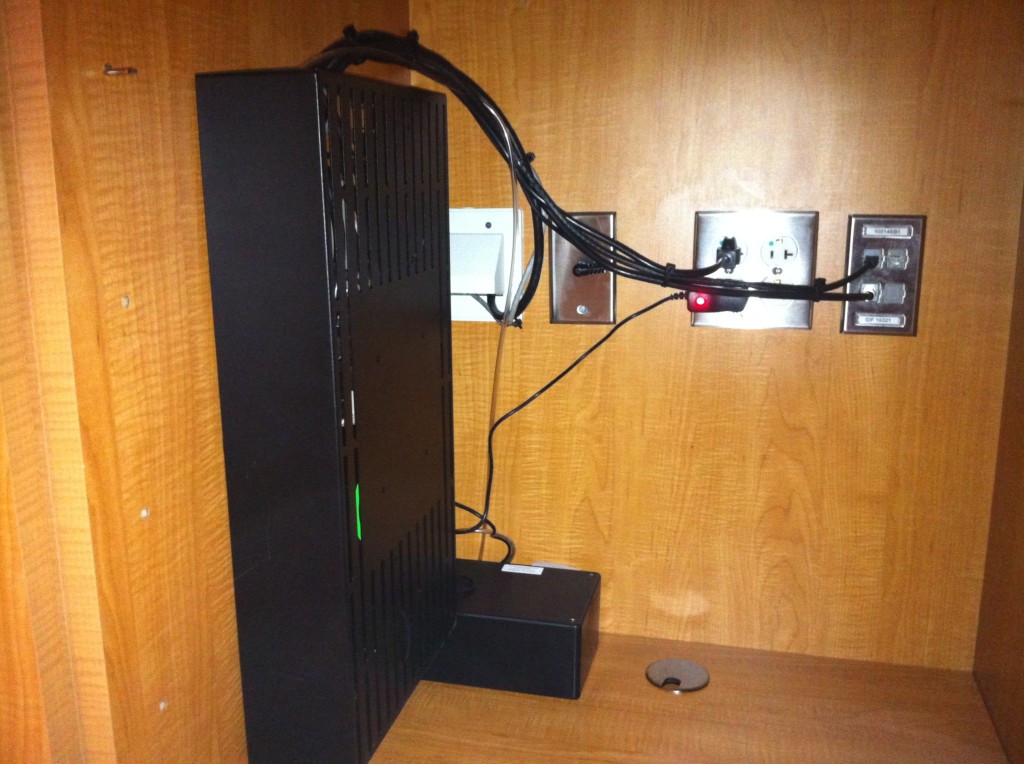
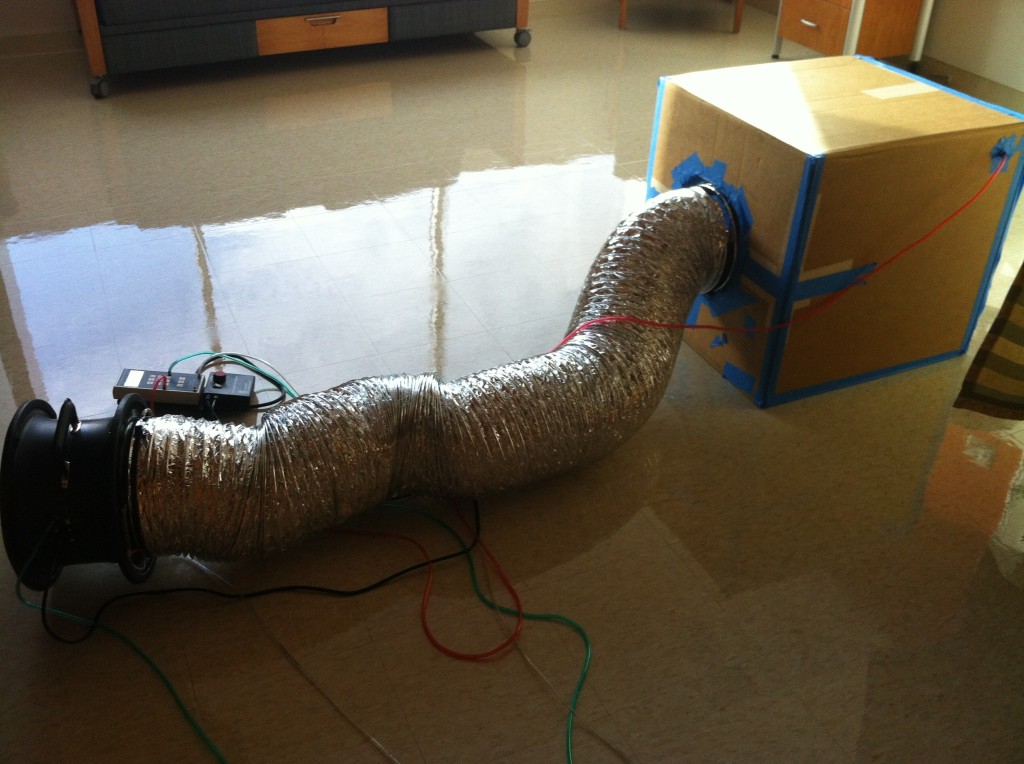
- Outdoor air ventilation rates (using CO2 measurements in the supply, return, and outdoor air streams of the air handling units in the mechanical rooms that serve each floor — we also measured airflow rates from the supply registers and return grilles in each patient room before the hospital opened)
This network of sensors therefore includes about 7 variables per room (times 10 rooms), plus 3 variables per nurse station (times 2 rooms), or about 76 variables in total. Each one of these is being measured and stored at 5 minute intervals over the course of a year, providing a whopping 105,000 time-series data points for each measured variable (12 data points per hour x 8760 hours per year). Multiply by the 76 or so variables we’re measuring and we’re talking 8 million or so individual data points!! (of course every once in a while a sensor goes missing, fails, or a battery dies, so the real number might be more like 7.5 million)
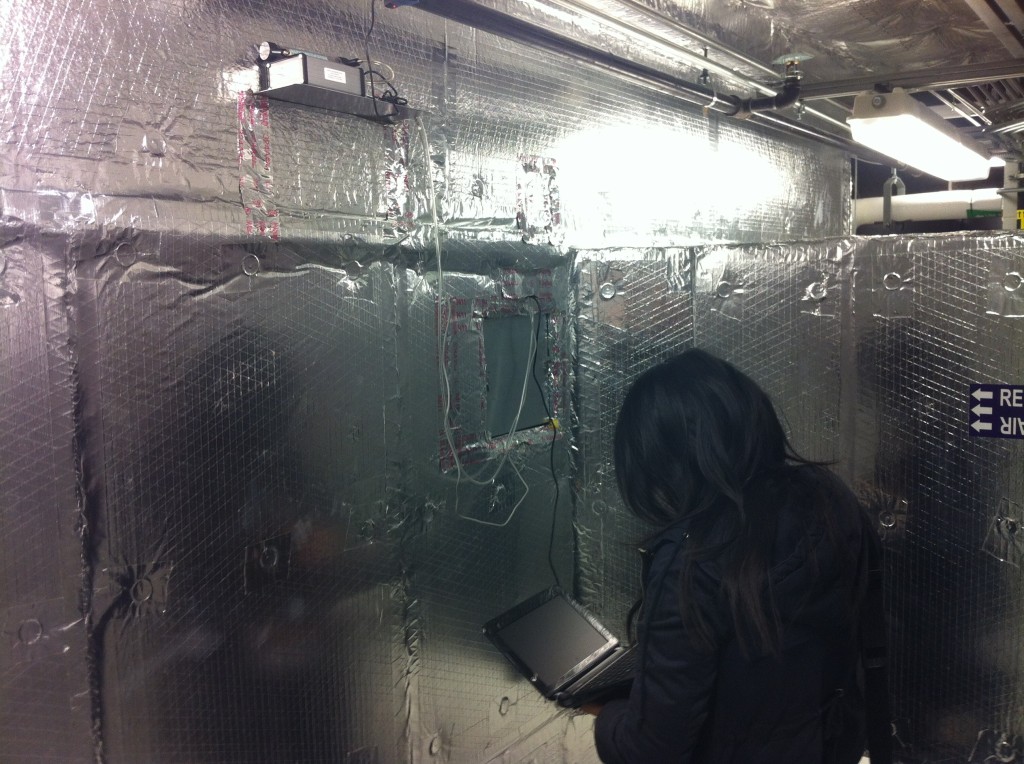

CO2 sensors in the mechanical rooms measure the fraction of outdoor air supplied by each HVAC system
Each of these sensors requires battery power and data storage, so maintaining them is a weekly job. Each week one or two of my students visits at the same time as Jack’s field team to download data and check up on sensors. Sometimes the whole team goes out to perform emergency sensor surgery when batteries fail and other issues arise. We’re also using HVAC filter media installed on the HVAC return grilles to passively sample airborne microbes (an interesting technique pioneered by Jeff Siegel).
And just to give you a sneak peak as to why we bother measuring all of these variables, it turns out that they can vary quite a lot from room to room. Check out some of these data from the project.
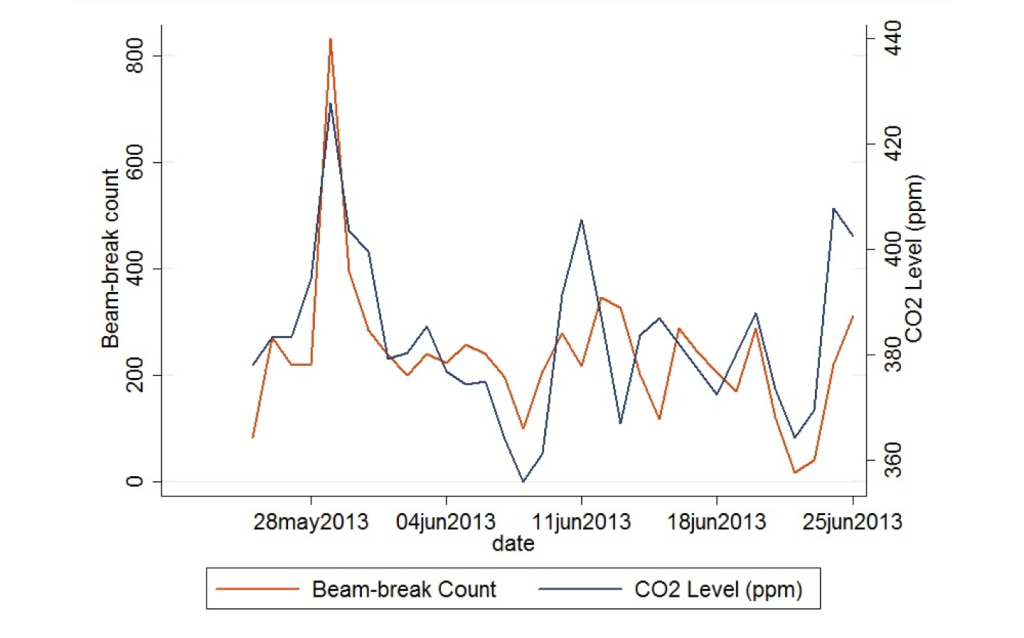
Daily beam break counts correlate well with daily mean CO2 concentrations, suggesting both can be used for human occupancy measurements
This is one of the largest building environmental characterizations for an indoor microbiology study of which I’m aware, and we’re very happy to be a part of it! Many thanks to Jeff Siegel for working on this experimental plan, Hal Levin and Jack Gilbert for helping to bring us on to this project, Paula Olsiewski at the Alfred P. Sloan Foundation, and my students Tiffanie Ramos, Parham Azimi, and Laurit Dide for their continuing work on data collection and sensor maintenance. I’ll be posting again in a few months with more results!
Some other links associated with the Hospital Microbiome Project:
April 2013 microBEnet blog post on the Hospital Microbiome Project by Hal Levin
January 2013 meeting report in Standards in Genomic Science
Filed under: Doing research | Experimental methods | Local to Chicago | Student awesomeness